Advanced_cate_estimation_approaches_molak_chap10
From Uplift Modeling to Counterfactual Explanations on a Public Experimental Dataset
This notebook recreates the main ideas and code patterns from Chapter 10 of Causal Inference and Machine Learning by Aleksander Molak, but on a different public dataset.
Instead of the Hillstrom email dataset used in the book, we will use the LaLonde job-training experiment, a classic randomized dataset that is publicly available. The goal is the same:
- verify randomization,
- fit several CATE / uplift estimators,
- compare compute cost,
- evaluate models with uplift-by-decile and expected response,
- extract confidence intervals,
- finish with a small section on counterfactual explanations.
Why this notebook is slightly adapted
The original chapter uses:
- a multi-treatment marketing dataset,
- and some code patterns tailored to that setup.
The LaLonde data is:
- a binary treatment experiment (
treatvscontrol), - with a continuous outcome (
re78, earnings in 1978).
So the notebook mirrors the chapter’s workflow, but simplifies a few formulas to the binary-treatment case.
Important: the final counterfactual section explains model behavior, not ground-truth causality. That is also the key message in Molak’s chapter.
# If you are running this notebook in a fresh environment, uncomment the next line.
!pip install -q econml dowhy dice-ml lightgbm rdatasets
import warnings
warnings.filterwarnings("ignore")
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from lightgbm import LGBMRegressor, LGBMClassifier
# EconML estimators
from econml.metalearners import SLearner, TLearner, XLearner
from econml.dr import DRLearner
from econml.dml import LinearDML, CausalForestDML
np.random.seed(42)
pd.set_option("display.max_columns", 200)
1. Load a different public dataset: LaLonde
The LaLonde dataset is a randomized job-training study and is widely used in causal inference.
We will download a public CSV mirror from the Rdatasets repository.
url = "https://raw.githubusercontent.com/vincentarelbundock/Rdatasets/master/csv/MatchIt/lalonde.csv"
df = pd.read_csv(url)
# Keep only the useful columns
df = df.drop(columns=["rownames"], errors="ignore")
# Treatment and outcome naming consistent with the chapter
df = df.rename(columns={"treat": "treatment", "re78": "outcome"})
df.head()
| treatment | age | educ | race | married | nodegree | re74 | re75 | outcome | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 37 | 11 | black | 1 | 1 | 0.0 | 0.0 | 9930.0460 |
| 1 | 1 | 22 | 9 | hispan | 0 | 1 | 0.0 | 0.0 | 3595.8940 |
| 2 | 1 | 30 | 12 | black | 0 | 0 | 0.0 | 0.0 | 24909.4500 |
| 3 | 1 | 27 | 11 | black | 0 | 1 | 0.0 | 0.0 | 7506.1460 |
| 4 | 1 | 33 | 8 | black | 0 | 1 | 0.0 | 0.0 | 289.7899 |
Data dictionary (quick version)
treatment: whether the person received job trainingoutcome: earnings in 1978age,educ: age and educationblack,hispan,married,nodegree: demographic indicatorsre74,re75: earnings in prior years
The experiment is randomized, which means treatment should not be predictable from covariates if randomization worked well.
df.describe(include="all").T
| count | unique | top | freq | mean | std | min | 25% | 50% | 75% | max | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| treatment | 614.0 | NaN | NaN | NaN | 0.301303 | 0.459198 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 |
| age | 614.0 | NaN | NaN | NaN | 27.363192 | 9.881187 | 16.0 | 20.0 | 25.0 | 32.0 | 55.0 |
| educ | 614.0 | NaN | NaN | NaN | 10.26873 | 2.628325 | 0.0 | 9.0 | 11.0 | 12.0 | 18.0 |
| race | 614 | 3 | white | 299 | NaN | NaN | NaN | NaN | NaN | NaN | NaN |
| married | 614.0 | NaN | NaN | NaN | 0.415309 | 0.493177 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 |
| nodegree | 614.0 | NaN | NaN | NaN | 0.630293 | 0.483119 | 0.0 | 0.0 | 1.0 | 1.0 | 1.0 |
| re74 | 614.0 | NaN | NaN | NaN | 4557.546569 | 6477.964479 | 0.0 | 0.0 | 1042.33 | 7888.49825 | 35040.07 |
| re75 | 614.0 | NaN | NaN | NaN | 2184.938207 | 3295.679043 | 0.0 | 0.0 | 601.5484 | 3248.9875 | 25142.24 |
| outcome | 614.0 | NaN | NaN | NaN | 6792.834483 | 7470.730792 | 0.0 | 238.283425 | 4759.0185 | 10893.5925 | 60307.93 |
2. Build feature, treatment, and outcome matrices
X = df.drop(columns=["treatment", "outcome"])
X = pd.get_dummies(X,drop_first=True)
T = df["treatment"].astype(int)
Y = df["outcome"].astype(float)
print("Rows:", len(df))
print("Treatment rate:", T.mean().round(4))
print("Average outcome:", Y.mean().round(2))
X.head()
Rows: 614
Treatment rate: 0.3013
Average outcome: 6792.83
| age | educ | married | nodegree | re74 | re75 | race_hispan | race_white | |
|---|---|---|---|---|---|---|---|---|
| 0 | 37 | 11 | 1 | 1 | 0.0 | 0.0 | False | False |
| 1 | 22 | 9 | 0 | 1 | 0.0 | 0.0 | True | False |
| 2 | 30 | 12 | 0 | 0 | 0.0 | 0.0 | False | False |
| 3 | 27 | 11 | 0 | 1 | 0.0 | 0.0 | False | False |
| 4 | 33 | 8 | 0 | 1 | 0.0 | 0.0 | False | False |
3. Randomization sanity check
Just like in the chapter, we first ask:
Can observed covariates predict treatment?
If treatment assignment is really random, a model should not do much better than naive guessing.
# Check marginal treatment distribution
treatment_dist = T.value_counts(normalize=True).sort_index()
treatment_dist
treatment
0 0.698697
1 0.301303
Name: proportion, dtype: float64
X_train_eda, X_test_eda, T_train_eda, T_test_eda = train_test_split(
X, T, test_size=0.5, random_state=42, stratify=T
)
clf_eda = LGBMClassifier(
n_estimators=100,
max_depth=4,
learning_rate=0.05,
verbosity=-1,
random_state=42
)
clf_eda.fit(X_train_eda, T_train_eda)
T_pred_eda = clf_eda.predict(X_test_eda)
eda_accuracy = accuracy_score(T_test_eda, T_pred_eda)
eda_accuracy
$\displaystyle 0.820846905537459$
For a binary treatment, the naive benchmark is roughly the larger class probability. Now we simulate what a random classifier would achieve if it only respected the treatment marginal.
p1 = T.mean()
n_test = len(T_test_eda)
random_scores = []
for _ in range(10000):
random_pred = np.random.binomial(1, p1, size=n_test)
random_scores.append((random_pred == T_test_eda.to_numpy()).mean())
ci_low, ci_high = np.quantile(random_scores, [0.025, 0.975])
print("Observed treatment-prediction accuracy :", round(eda_accuracy, 4))
print("95% empirical random-accuracy interval :", (round(ci_low, 4), round(ci_high, 4)))
Observed treatment-prediction accuracy : 0.8208
95% empirical random-accuracy interval : (np.float64(0.5277), np.float64(0.6319))
plt.figure(figsize=(8, 4.5))
plt.hist(random_scores, bins=40, alpha=0.8)
plt.axvline(eda_accuracy, linestyle="--", linewidth=2, label="Model accuracy")
plt.title("Randomization check: model accuracy vs random baseline")
plt.xlabel("Accuracy")
plt.ylabel("Count")
plt.legend()
plt.show()
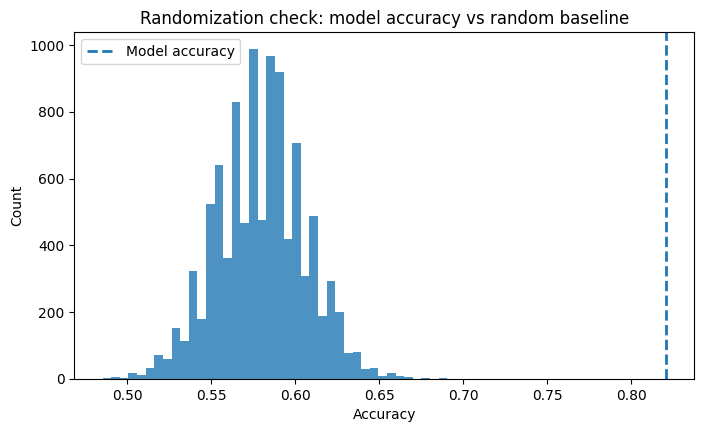
If the model accuracy falls near the random baseline, that is good news: the observed covariates do not strongly predict treatment, which is what we hope to see in a randomized experiment.
4. Train / test split for uplift modeling
Following the chapter, we create a separate train and test split. Because experimental datasets can still be small in terms of effective signal, it is useful to keep a large test set.
X_train, X_test, y_train, y_test, T_train, T_test = train_test_split(
X, Y, T, test_size=0.5, random_state=42, stratify=T
)
print("Train rows:", len(X_train))
print("Test rows :", len(X_test))
print("Treatment rate train:", T_train.mean().round(4))
print("Treatment rate test :", T_test.mean().round(4))
Train rows: 307
Test rows : 307
Treatment rate train: 0.3029
Treatment rate test : 0.2997
5. Helper functions and model definitions
We now recreate the same family of estimators used in the chapter:
- S-Learner
- T-Learner
- X-Learner
- DR-Learner
- Linear DML
- Causal Forest DML
For consistency, we use LightGBM as the main base learner, just like the book often does.
def create_regressor():
return LGBMRegressor(
n_estimators=200,
max_depth=4,
learning_rate=0.05,
subsample=0.9,
colsample_bytree=0.9,
verbosity=-1,
random_state=42
)
def create_classifier():
return LGBMClassifier(
n_estimators=200,
max_depth=4,
learning_rate=0.05,
subsample=0.9,
colsample_bytree=0.9,
verbosity=-1,
random_state=42
)
s_learner = SLearner(overall_model=create_regressor())
t_learner = TLearner(models=[create_regressor(), create_regressor()])
x_learner = XLearner(
models=[create_regressor(), create_regressor()],
cate_models=[create_regressor(), create_regressor()],
)
dr_learner = DRLearner(
model_propensity=LogisticRegression(max_iter=2000),
model_regression=create_regressor(),
model_final=create_regressor(),
cv=5,
)
linear_dml = LinearDML(
model_y=create_regressor(),
model_t=create_classifier(),
discrete_treatment=True,
cv=5,
random_state=42,
)
causal_forest = CausalForestDML(
model_y=create_regressor(),
model_t=create_classifier(),
discrete_treatment=True,
cv=5,
random_state=42,
)
models = {
"SLearner": s_learner,
"TLearner": t_learner,
"XLearner": x_learner,
"DRLearner": dr_learner,
"LinearDML": linear_dml,
"CausalForestDML": causal_forest,
}
6. Fit all models and compare training time
This mirrors the timing comparison in the chapter. Exact times will vary by machine, but relative ordering is the main idea.
fit_times = {}
for model_name, model in models.items():
start = time.time()
if model_name in {"LinearDML", "CausalForestDML"}:
model.fit(Y=y_train, T=T_train, X=X_train)
else:
model.fit(Y=y_train, T=T_train, X=X_train)
stop = time.time()
fit_times[model_name] = stop - start
print(f"{model_name:<16} fitted in {fit_times[model_name]:.3f} seconds")
SLearner fitted in 0.052 seconds
TLearner fitted in 0.088 seconds
XLearner fitted in 0.129 seconds
DRLearner fitted in 2.457 seconds
LinearDML fitted in 0.371 seconds
CausalForestDML fitted in 0.583 seconds
timing_df = pd.DataFrame({
"Model": list(fit_times.keys()),
"TimeSeconds": list(fit_times.values())
}).sort_values("TimeSeconds")
baseline = timing_df["TimeSeconds"].min()
timing_df["RelativeToFastest"] = (timing_df["TimeSeconds"] / baseline).round(1)
timing_df
| Model | TimeSeconds | RelativeToFastest | |
|---|---|---|---|
| 0 | SLearner | 0.051677 | 1.0 |
| 1 | TLearner | 0.087885 | 1.7 |
| 2 | XLearner | 0.129461 | 2.5 |
| 4 | LinearDML | 0.370959 | 7.2 |
| 5 | CausalForestDML | 0.582778 | 11.3 |
| 3 | DRLearner | 2.456517 | 47.5 |
plt.figure(figsize=(9, 4.5))
plt.bar(timing_df["Model"], timing_df["RelativeToFastest"])
plt.xticks(rotation=30, ha="right")
plt.ylabel("Relative training time")
plt.title("Relative compute cost across CATE estimators")
plt.show()
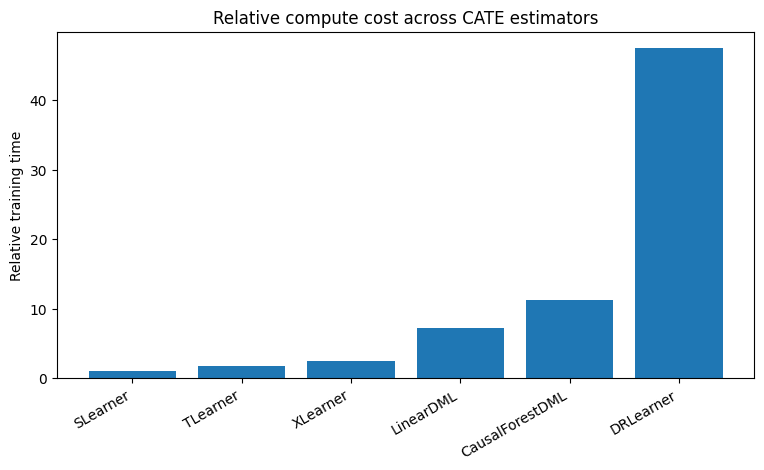
7. Get CATE / uplift predictions
For a binary treatment experiment, the predicted uplift is simply the estimated effect of going from control (T0=0) to treatment (T1=1).
def cate_predict(model, X_data):
return model.effect(X_data)
cate_train = {name: cate_predict(model, X_train) for name, model in models.items()}
cate_test = {name: cate_predict(model, X_test) for name, model in models.items()}
pd.DataFrame({k: np.asarray(v).ravel()[:5] for k, v in cate_test.items()})
| SLearner | TLearner | XLearner | DRLearner | LinearDML | CausalForestDML | |
|---|---|---|---|---|---|---|
| 0 | 66.838847 | 6255.915004 | 3166.608390 | 3546.645036 | 5768.924299 | 3262.998457 |
| 1 | -137.971741 | -4166.883389 | -616.743227 | -4107.529289 | 8828.882399 | 884.823406 |
| 2 | 1596.118279 | 2900.016716 | 474.007860 | -5306.252895 | 3452.218217 | 1884.682510 |
| 3 | 539.519164 | 294.686420 | 1326.560882 | 101.825536 | 1065.778789 | 950.812086 |
| 4 | 494.817338 | 4926.474853 | 2585.430688 | 6747.790039 | -353.910100 | 207.545285 |
8. Uplift by decile
This is one of the chapter’s main ideas.
Intuition
- Score each unit by predicted uplift.
- Sort from highest predicted uplift to lowest.
- Split into 10 bins (deciles).
- Inside each decile, estimate the observed uplift: Observed uplift = E[Y | T = 1] − E[Y | T = 0]
- A good model should show higher observed uplift in top deciles than in lower deciles.
def uplift_by_decile(y, t, uplift, n_bins=10):
temp = pd.DataFrame({
"y": np.asarray(y),
"t": np.asarray(t).astype(int),
"uplift": np.asarray(uplift).ravel()
}).sort_values("uplift", ascending=False).reset_index(drop=True)
temp["decile"] = pd.qcut(
np.arange(len(temp)),
q=n_bins,
labels=False,
duplicates="drop"
)
rows = []
for decile, group in temp.groupby("decile"):
treated = group.loc[group["t"] == 1, "y"]
control = group.loc[group["t"] == 0, "y"]
uplift_obs = treated.mean() - control.mean() if len(treated) and len(control) else np.nan
rows.append({
"decile": int(decile),
"n": len(group),
"treated_n": len(treated),
"control_n": len(control),
"observed_uplift": uplift_obs
})
return pd.DataFrame(rows)
def plot_uplift_by_decile(decile_df, title):
plt.figure(figsize=(8, 4.5))
plt.bar(decile_df["decile"], decile_df["observed_uplift"])
plt.axhline(0, linewidth=1)
plt.title(title)
plt.xlabel("Decile (0 = highest predicted uplift)")
plt.ylabel("Observed uplift")
plt.show()
# Example: one model on train and test
example_train = uplift_by_decile(y_train, T_train, cate_train["XLearner"])
example_test = uplift_by_decile(y_test, T_test, cate_test["XLearner"])
plot_uplift_by_decile(example_train, "XLearner — uplift by decile (train)")
plot_uplift_by_decile(example_test, "XLearner — uplift by decile (test)")
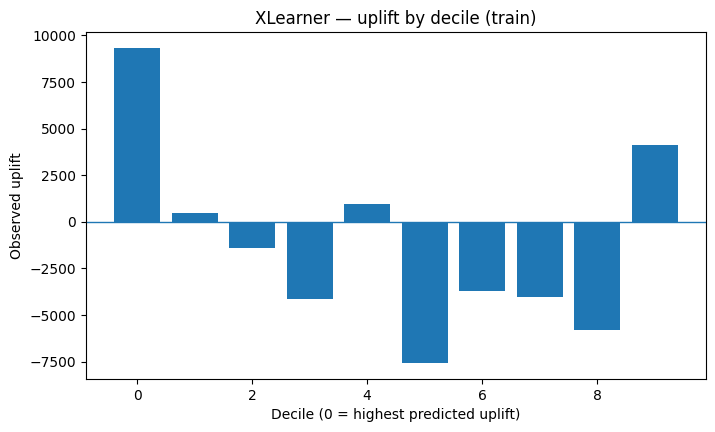
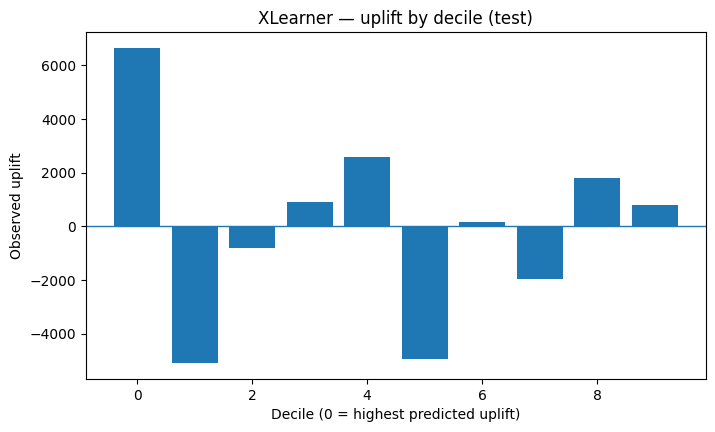
Compare all models
fig, axes = plt.subplots(len(models), 2, figsize=(12, 3.4 * len(models)))
for row_idx, (name, _) in enumerate(models.items()):
train_df = uplift_by_decile(y_train, T_train, cate_train[name])
test_df = uplift_by_decile(y_test, T_test, cate_test[name])
axes[row_idx, 0].bar(train_df["decile"], train_df["observed_uplift"])
axes[row_idx, 0].axhline(0, linewidth=1)
axes[row_idx, 0].set_title(f"{name} — Train")
axes[row_idx, 0].set_xlabel("Decile")
axes[row_idx, 0].set_ylabel("Obs uplift")
axes[row_idx, 1].bar(test_df["decile"], test_df["observed_uplift"])
axes[row_idx, 1].axhline(0, linewidth=1)
axes[row_idx, 1].set_title(f"{name} — Test")
axes[row_idx, 1].set_xlabel("Decile")
axes[row_idx, 1].set_ylabel("Obs uplift")
plt.tight_layout()
plt.show()
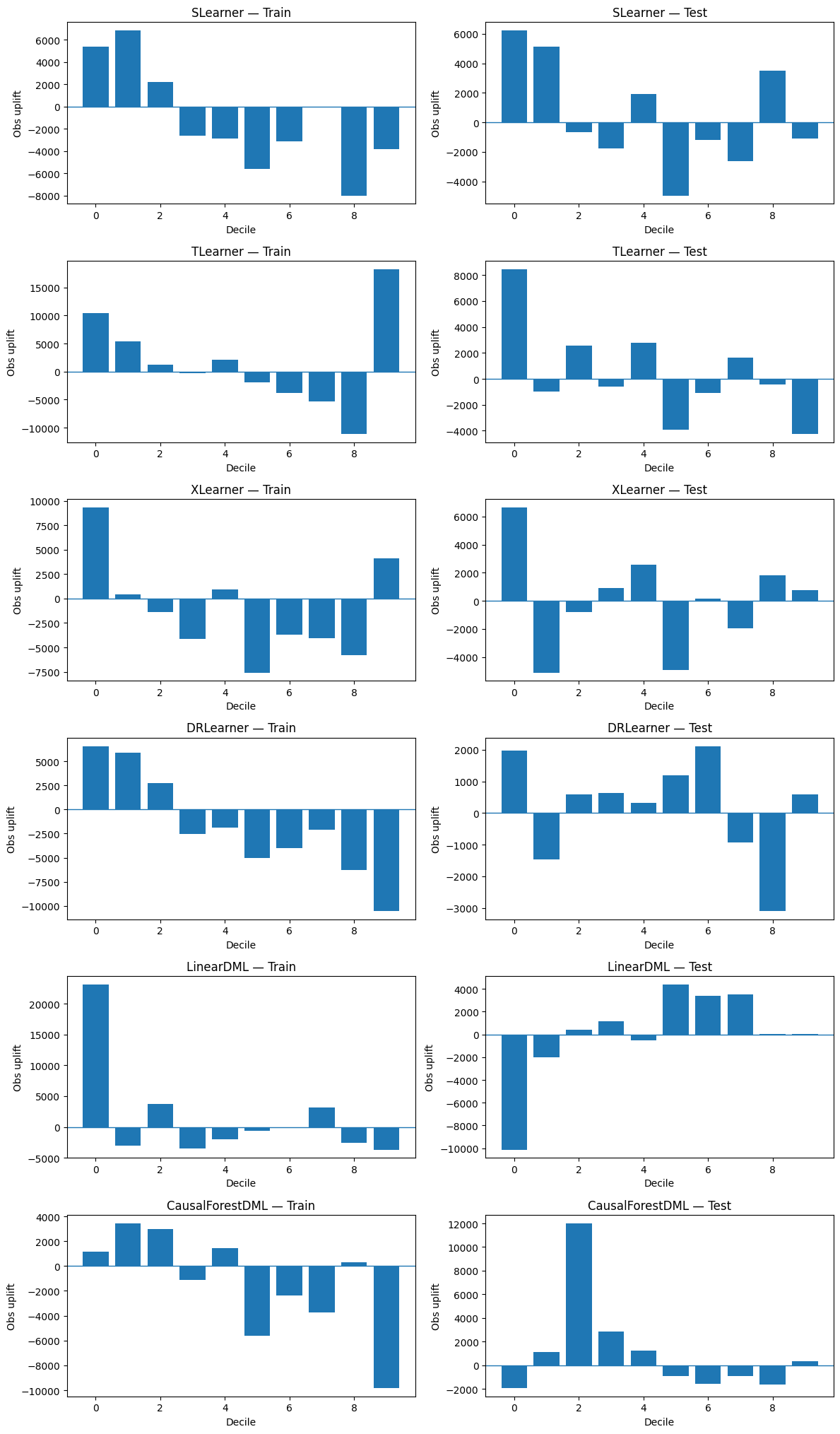
9. Expected response (binary-treatment version)
In the chapter, expected response is used as a more decision-oriented metric.
Intuition
For each person:
- the model recommends treatment if predicted uplift is positive,
- otherwise the model recommends control.
Because we only observe the outcome under the actually assigned treatment, we estimate how good the policy is with an inverse-propensity-weighted policy value:
| Expected Response = E[ Y × I(T = a(X)) / P(T | X) ] |
Where I(T = a(X)) is equal to 1, when treatment is actually assigned, else 0. In a randomized experiment with roughly constant assignment probability, this becomes straightforward to compute.
def expected_response_binary(y, t, uplift_scores, p_treat=None):
y = np.asarray(y)
t = np.asarray(t).astype(int)
uplift_scores = np.asarray(uplift_scores).ravel()
if p_treat is None:
p_treat = t.mean()
p_control = 1 - p_treat
recommended_treatment = (uplift_scores > 0).astype(int)
weight = np.where(recommended_treatment == 1, 1 / p_treat, 1 / p_control)
observed_if_followed = (recommended_treatment == t).astype(int)
return np.mean(y * observed_if_followed * weight)
metric_rows = []
for name in models:
train_er = expected_response_binary(y_train, T_train, cate_train[name], p_treat=T_train.mean())
test_er = expected_response_binary(y_test, T_test, cate_test[name], p_treat=T_test.mean())
metric_rows.append({
"Model": name,
"ExpectedResponse_Train": train_er,
"ExpectedResponse_Test": test_er
})
expected_response_df = pd.DataFrame(metric_rows).sort_values("ExpectedResponse_Test", ascending=False)
expected_response_df
| Model | ExpectedResponse_Train | ExpectedResponse_Test | |
|---|---|---|---|
| 0 | SLearner | 10084.517560 | 9177.923448 |
| 1 | TLearner | 10047.760508 | 8225.655137 |
| 2 | XLearner | 9822.502760 | 8089.190804 |
| 5 | CausalForestDML | 8044.869124 | 7214.912195 |
| 3 | DRLearner | 8838.831954 | 5989.903949 |
| 4 | LinearDML | 5440.521197 | 4828.437861 |
plt.figure(figsize=(9, 4.5))
plt.bar(expected_response_df["Model"], expected_response_df["ExpectedResponse_Test"])
plt.xticks(rotation=30, ha="right")
plt.ylabel("Expected response (test)")
plt.title("Policy value comparison on the test set")
plt.show()
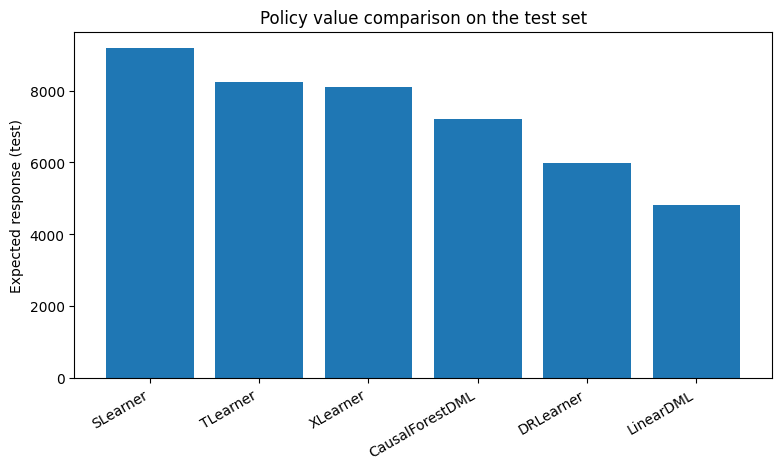
10. Confidence intervals with Linear DML
One nice feature highlighted in the chapter is that LinearDML can provide confidence intervals directly.
lb, ub = models["LinearDML"].effect_interval(X_test, T0=0, T1=1, alpha=0.05)
print("Lower bounds (first 5):", lb[:5])
print("Upper bounds (first 5):", ub[:5])
Lower bounds (first 5): [ -288.91205216 2458.50208897 -2216.77656149 -4170.36233339
-3868.06794743]
Upper bounds (first 5): [11826.76064953 15199.26270957 9121.21299571 6301.91991069
3160.24774699]
intervals = np.column_stack([lb, ub])
contains_zero = np.mean(np.sign(intervals[:, 0]) != np.sign(intervals[:, 1]))
print("Fraction of test observations whose 95% CI contains 0:", round(float(contains_zero), 4))
Fraction of test observations whose 95% CI contains 0: 0.8534
plt.figure(figsize=(8, 4.5))
sample_idx = np.arange(min(50, len(lb)))
plt.errorbar(
sample_idx,
cate_test["LinearDML"][:len(sample_idx)],
yerr=[
cate_test["LinearDML"][:len(sample_idx)] - lb[:len(sample_idx)],
ub[:len(sample_idx)] - cate_test["LinearDML"][:len(sample_idx)]
],
fmt='o'
)
plt.axhline(0, linewidth=1)
plt.title("LinearDML: point estimates with 95% confidence intervals")
plt.xlabel("Sample index")
plt.ylabel("Estimated treatment effect")
plt.show()
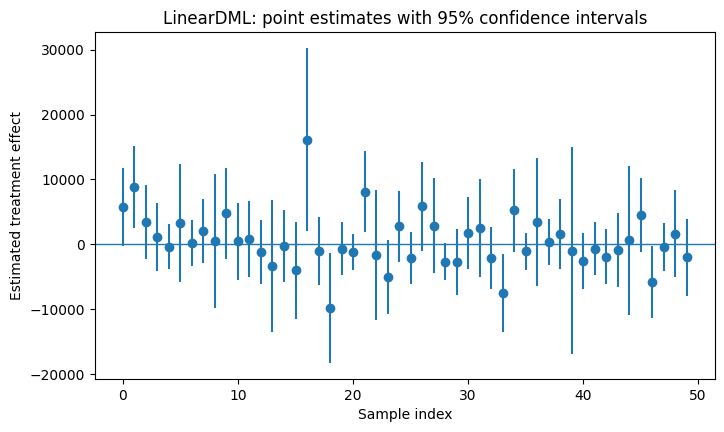
Optional targeting rule
A very practical rule is:
- only target people whose estimated uplift is positive and
- whose interval does not include zero.
That gives you a more conservative treatment policy.
conservative_treat = (
(np.asarray(cate_test["LinearDML"]).ravel() > 0) &
(lb > 0)
).astype(int)
pd.Series(conservative_treat).value_counts(normalize=True).rename("share")
0 0.879479
1 0.120521
Name: share, dtype: float64
11. A compact comparison table
This is not the exact table from the book, but it serves the same purpose: bring together practical aspects you might care about.
comparison = expected_response_df.merge(
timing_df[["Model", "RelativeToFastest"]],
on="Model",
how="left"
).rename(columns={"RelativeToFastest": "RelativeComputeCost"})
comparison
| Model | ExpectedResponse_Train | ExpectedResponse_Test | RelativeComputeCost | |
|---|---|---|---|---|
| 0 | SLearner | 10084.517560 | 9177.923448 | 1.0 |
| 1 | TLearner | 10047.760508 | 8225.655137 | 1.7 |
| 2 | XLearner | 9822.502760 | 8089.190804 | 2.5 |
| 3 | CausalForestDML | 8044.869124 | 7214.912195 | 11.3 |
| 4 | DRLearner | 8838.831954 | 5989.903949 | 47.5 |
| 5 | LinearDML | 5440.521197 | 4828.437861 | 7.2 |
12. Extra: counterfactual explanations
The chapter ends with a short section on counterfactual explanations.
Key idea
This is not about estimating the true causal effect in the world. It is about asking:
What small change to the input would flip the model’s decision?
To demonstrate this in a way that is close to the chapter, we will:
- take the estimated uplift from
LinearDML, - convert it into a simple recommendation label:
1if the model recommends treatment,0if the model recommends control,
- use DiCE to find feature changes that would flip the recommendation.
This explains the decision policy, not the true data-generating mechanism.
# Create a recommendation label from the LinearDML uplift estimates
train_recommend = (np.asarray(cate_train["LinearDML"]).ravel() > 0).astype(int)
test_recommend = (np.asarray(cate_test["LinearDML"]).ravel() > 0).astype(int)
recommend_train_df = X_train.copy()
recommend_train_df["recommend_treatment"] = train_recommend
recommend_test_df = X_test.copy()
recommend_test_df["recommend_treatment"] = test_recommend
recommend_train_df.head()
| age | educ | married | nodegree | re74 | re75 | race_hispan | race_white | recommend_treatment | |
|---|---|---|---|---|---|---|---|---|---|
| 19 | 26 | 12 | 0 | 0 | 0.000 | 0.000 | False | False | 0 |
| 52 | 18 | 11 | 0 | 1 | 0.000 | 0.000 | False | False | 1 |
| 296 | 28 | 13 | 0 | 0 | 5260.631 | 3790.113 | False | True | 1 |
| 37 | 23 | 12 | 1 | 0 | 0.000 | 0.000 | False | False | 1 |
| 369 | 18 | 10 | 0 | 1 | 0.000 | 1491.339 | False | True | 0 |
# Train a small interpretable recommendation model for DiCE
recommendation_model = LogisticRegression(max_iter=5000)
recommendation_model.fit(X_train, train_recommend)
print("Share recommended for treatment in train:", train_recommend.mean().round(3))
print("Share recommended for treatment in test :", test_recommend.mean().round(3))
Share recommended for treatment in train: 0.638
Share recommended for treatment in test : 0.642
# DiCE setup
import dice_ml
from dice_ml import Dice
dice_data = dice_ml.Data(
dataframe=recommend_train_df,
continuous_features=["age", "educ", "re74", "re75"],
outcome_name="recommend_treatment",
)
dice_model = dice_ml.Model(model=recommendation_model, backend="sklearn", model_type="classifier")
dice = Dice(dice_data, dice_model, method="random")
# Pick one person the policy currently does NOT recommend for treatment
candidate_pool = recommend_test_df.copy()
candidate_pool["pred"] = recommendation_model.predict(X_test)
query_idx = candidate_pool.index[candidate_pool["pred"] == 0][0]
query_instance = X_test.loc[[query_idx]]
query_instance
| age | educ | married | nodegree | re74 | re75 | race_hispan | race_white | |
|---|---|---|---|---|---|---|---|---|
| 372 | 17 | 10 | 0 | 1 | 0.0 | 1453.742 | True | False |
cf = dice.generate_counterfactuals(
query_instance,
total_CFs=3,
desired_class="opposite",
features_to_vary=["age", "educ", "re74", "re75", "married", "nodegree"]
)
cf.visualize_as_dataframe(show_only_changes=True)
100%|████████████████████████████████████████████████████████████████████████████████████| 1/1 [00:00<00:00, 5.55it/s]
Query instance (original outcome : 0)
| age | educ | married | nodegree | re74 | re75 | race_hispan | race_white | recommend_treatment | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 17 | 10 | 0 | 1 | 0.0 | 1453.741943 | True | True | 0 |
Diverse Counterfactual set (new outcome: 1)
| age | educ | married | nodegree | re74 | re75 | race_hispan | race_white | recommend_treatment | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | - | - | - | - | 2584.0 | - | - | False | 1 |
| 1 | 34 | - | - | - | - | - | - | False | 1 |
| 2 | - | - | - | - | 17788.4 | - | - | False | 1 |
How to read the counterfactuals
Each row says something like:
“If this person’s features changed in these small ways, the recommendation model would flip from ‘do not treat’ to ‘treat’.”
Again, this is an explanation of the policy model, not a guarantee about the real world. That is exactly the spirit of the final section in Molak’s chapter.
13. Practical takeaways
What this notebook reproduced from the chapter
- randomized-experiment sanity checks,
- S / T / X / DR / LinearDML / CausalForestDML estimators,
- fit-time comparison,
- uplift-by-decile evaluation,
- expected response / policy value,
- confidence intervals,
- counterfactual explanations.
What changed from the chapter
- We used a different public dataset (LaLonde, not Hillstrom).
- The setup is binary treatment rather than multi-treatment.
- The final counterfactual section explains the recommendation model, which is the cleanest way to mirror the chapter on this dataset.